by

One of the pillars of science is the idea that experimental results can be replicated. If they cannot be reproduced, what if the findings of an experiment were due just to chance? Over the last two decades, a growing chorus of scientists has raised concerns about the “reproducibility crisis,” in which many published research findings can’t be independently validated, calling into question the rigor of contemporary science.
Two years ago, a group led by Konrad Kording, a Penn Integrates Knowledge Professor in Bioengineering and Neuroscience, founded the Community For Rigor (C4R) to build a grassroots movement to improve the rigor of scientific research.
Supported by a grant from the National Institutes of Health (NIH) and partners at Harvard, Duquesne, Smith College and Johns Hopkins, among other institutions, C4R creates educational materials that teach the principles of rigorous research, from data collection to pre-registration of results. “Everyone has done wrong things,” says Kording. “We’re all making these mistakes and we need to be able to talk about it.”
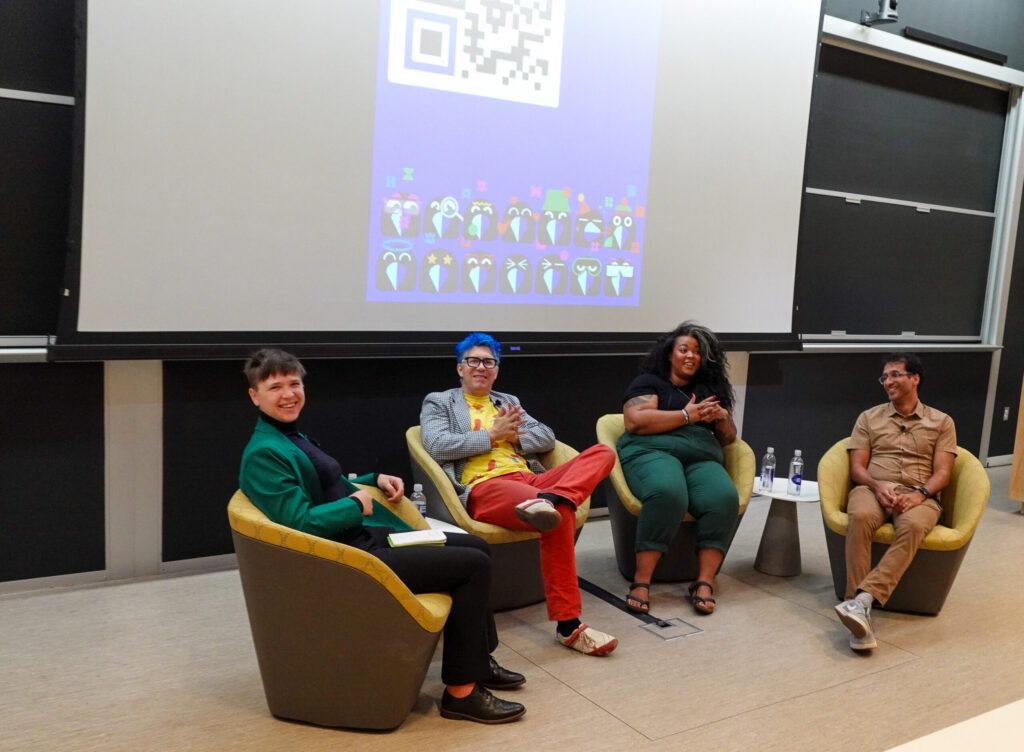
Last month, Kording appeared on In Plain English, a podcast devoted to making science more accessible, alongside Kaela Singleton, the co-founder and President of Black in Neuro; Arjun Raj, Professor in Bioengineering in Penn Engineering and in Genetics in Penn Medicine; and Jamie Moffa, a physician-scientist in training at Washington University in St. Louis, to discuss scientific rigor, including actionable strategies for students and faculty alike.
The conversation touched on everything from successfully managing the reams of data produced by experiments to the power of community to drive cultural change, as well as the difficulty of filtering useful feedback from the noise of social media. “I hope we can get to a point where people feel comfortable sharing what’s working and what’s not working,” says Raj.
Listen to the episode here.
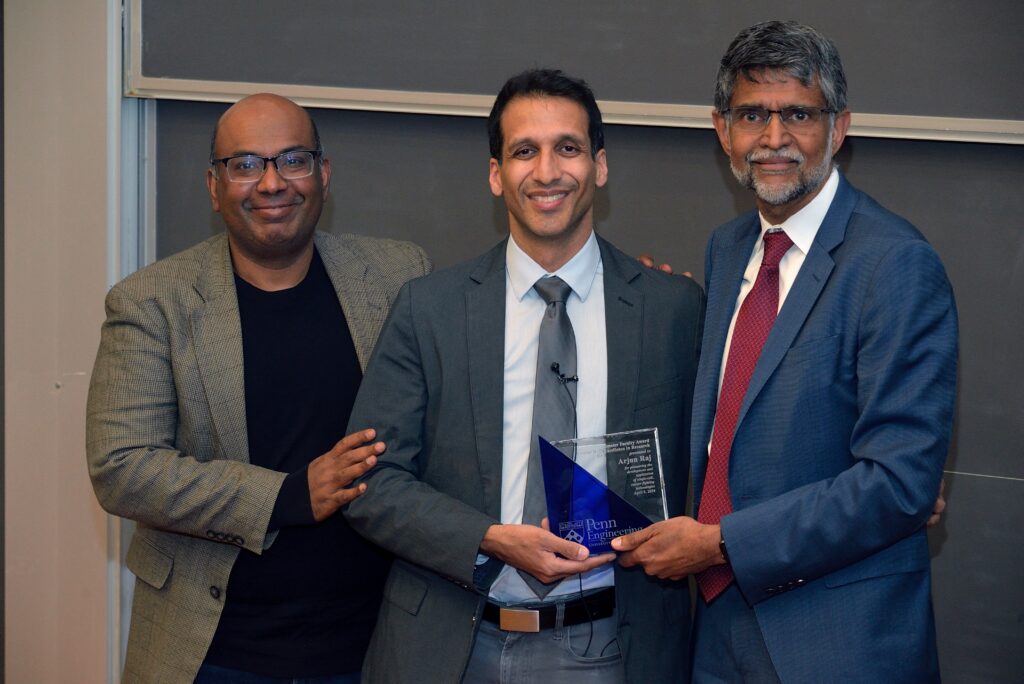


 A research team led by engineers at the
A research team led by engineers at the 

 Shawn Kang, BSE/MSE, Bioengineering (’22)
Shawn Kang, BSE/MSE, Bioengineering (’22)



