Brianna Leung, a rising senior majoring in Bioengineering and minoring in Neuroscience and Healthcare Management at the University of Pennsylvania, led a diverse team of student scientists and engineers to resounding success at the 2024 Cornell Health Tech Hackathon, where the team won the $3,000 Grand Prize.
Held in March 2024 on Cornell’s campus in New York City, the event brought together students from 29 different universities for a weekend of finding “hacks” to patient wellness and healthcare issues inspired by the theme of “patient safety.”

Leung serves as President of Penn Assistive Devices and Prosthetic Technologies (ADAPT), a medical-device project club whose members pursue personal projects, community partnerships and national design competitions. Penn ADAPT’s activities range from designing, building and improving assistive medical devices for conditions such as cerebral palsy and limb loss, to community engagement activities like their semesterly 3D-printed pancake sale.
In her role, Leung has increased the program’s hackathon participation to give club members greater exposure to fast-paced, competition-based design. She also leads the HMS School project, which develops and manufactures switch interfaces for children with cerebral palsy, enabling these students to interact with computers.
Leung’s passion for medical devices extends to her academic research. As a member of the robotics lab of Cynthia Sung, Gabel Family Term Assistant Professor in Mechanical Engineering and Applied Mechanics, Computer and Information Science, and Electrical and Systems Engineering, Leung characterizes origami patterns for energy-saving applications in the heart and in facial reconstruction. Leung has also served as Vice President External for the Penn Lions and Vice President of Member Engagement for the Wharton Undergraduate Healthcare Club, and belongs to the Phi Gamma Nu professional business fraternity.
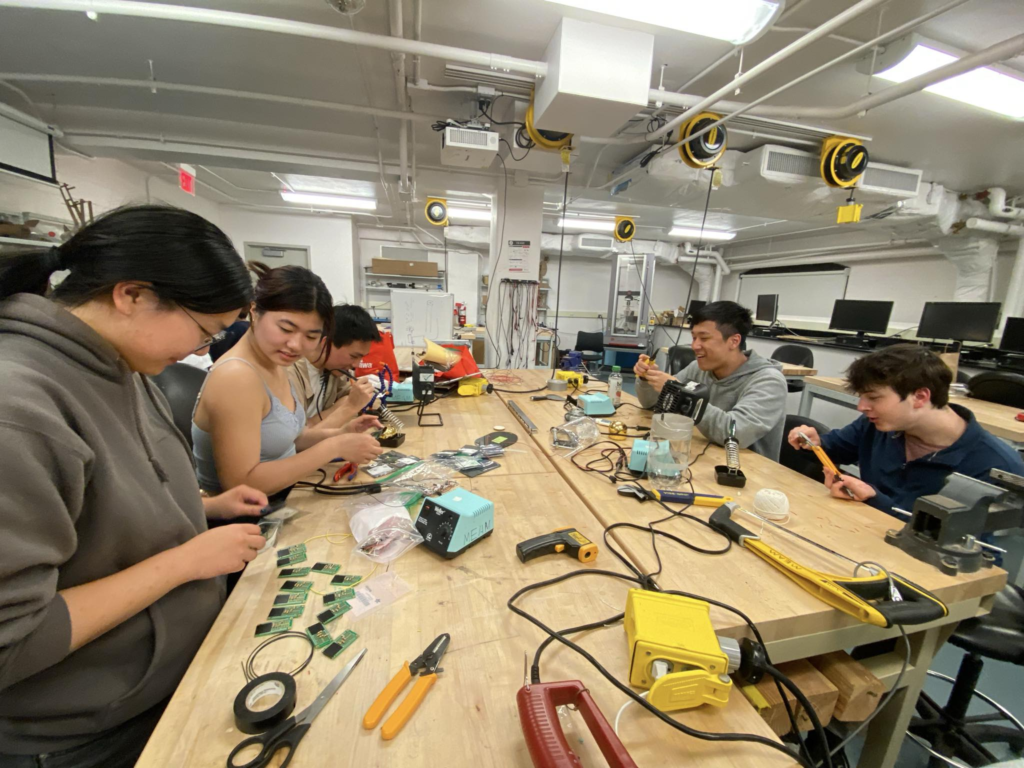

For the Cornell Hackathon, Leung’s team developed a prototype for Current Care, a closed-loop device to prevent pressure ulcers through electrical muscle stimulation. Pressure ulcers, often called bed sores, result from prolonged pressure, which often occurs during extended hospitalization or in patients who are bedridden. This condition is exacerbated by understaffing and strained resources, and can create an extra burden on hospitals, patients and healthcare workers. The U.S. Department of Health and Human Services estimates that pressure ulcers cost the U.S. healthcare system approximately $9.1 billion to $11.6 billion per year.
Current Care is designed to deliver electrical stimulation, which increases blood flow to affected body parts. Conceptualizing and designing complex devices on short notice is the nature of a hackathon, so the team focused their efforts on creating proof-of-concept prototypes for all the different sensors required for the device, as well as providing the judges with on-screen read-outs to demonstrate the logic and hypothetical inputs for the device.
For their design, the team was awarded the $3,000 Grand Prize in the Cornell Hackathon. In addition to Leung, the team consisted of Johnson Liu (Cornell ECE & MSE’26); Antranig Baghdassarian (Cornell BME’27); Andrew Lee (Weill Cornell M.D.’25); Leah Lackey (Cornell ECE Ph.D.’28); and Justin Liu (Northeastern CS’27).
In choosing a project, Leung was inspired by her late grandmother’s experiences. “My role on the team largely consisted of coordinating and leading aspects of its development as needed. I also ultimately presented our idea to the judges,” she says. “This was actually all of my teammates’ first hackathon, so it was really exciting to serve a new role (considering it was actually only my second hackathon!). I had a lot of fun working with them, and we have actually been meeting regularly since the event to continue to work on the project. We had a range of expertise and experience on our team, and I deeply appreciate their hard work and enthusiasm for a project that means so much to me.”
Having found success at the Cornell hackathon, the team is discussing next steps for Current Care. “Our team is still very motivated to continue working on the project, and we’ve been speaking with professors across all of our schools to discuss feasibility and design plans moving forward,” says Leung.
Several other projects developed by Penn ADAPT members were recognized in the Cornell Hackathon:

- Claire Zhang, a sophomore studying Bioengineering and Biology in the VIPER program, was a member and presenter for team CEDAR (winner of Most Innovative/2nd Place), a portable ultrasound imaging device used to monitor carotid artery stenosis development in rural areas.
- Natey Kim, a sophomore in Bioengineering, was a member and presenter for team HMSS (finalist), a low-cost digital solution for forecasting infections in hospitals.
- Rebecca Wang, a sophomore in Bioengineering and Social Chair of Penn ADAPT, was a member of Team Femnostics (winner of Most Market Ready/4th Place) which developed QuickSense, an all-in-one diagnostic tool that streamlines testing for a handful of the most common vaginal disease infections simultaneously.
- Mariam Rizvi, a sophomore in Computational Biology, was a member of team IPVision (winner of Most Potential Impact/5th Place), an application programming interface (or API) that integrates into electronic health records such as Epic, leveraging AI to detect intimate partner violence cases and provide personalized treatment in acute-care settings.
- Suhani Patel and Dwight Koyner worked with team RealAIs, which developed a full-stack multi-platform application using React Native and Vertex AI on the Google Cloud Platform (GCP). Patel, a sophomore double majoring in Bioengineering and Computer and Information Science in Penn Engineering, serves as ADAPT’s treasurer, while Koyner is a first-year M&T student studying Business and Systems Engineering in Penn Engineering and Wharton.




Learn more about Penn ADAPT here and follow their Instagram.
Read more about the 2024 Cornell Tech Hackathon in the Cornell Chronicle.