An Immune Cell Atlas
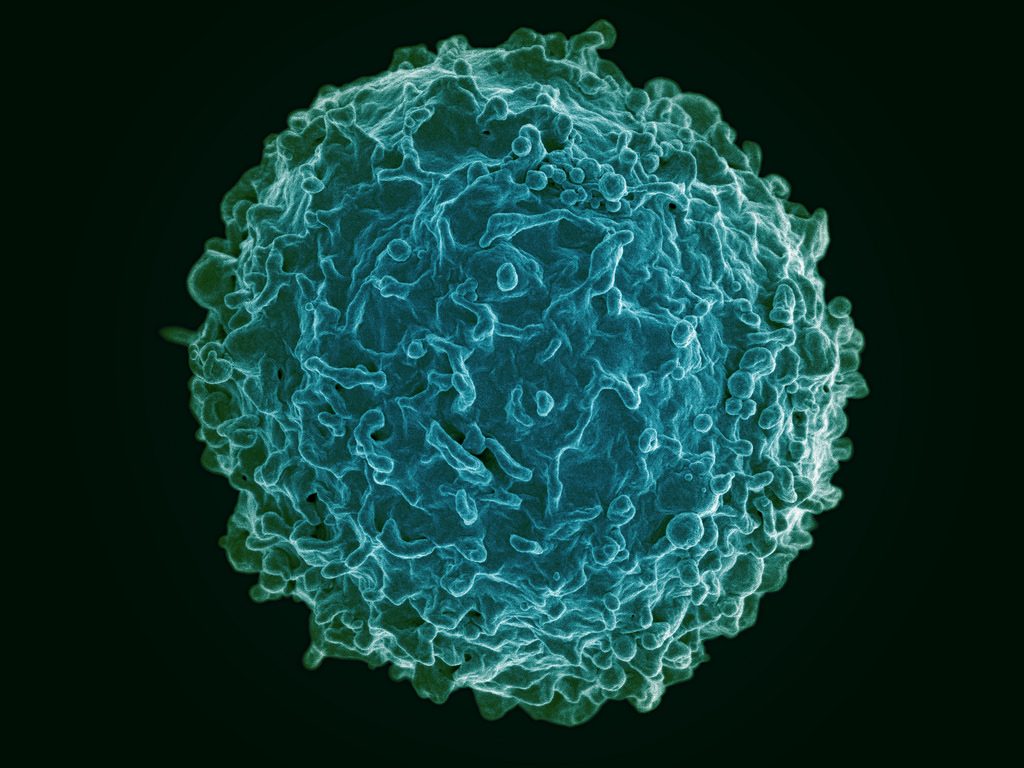
The human immune system deploys a variety of cells to counteract pathogens when they enter the body. B cells are a type of white blood cell specific to particular pathogens, and they form part of the adaptive immune system. As these cells develop, the cells with the strongest reactions to antigens are favored over others. This process is called clonal selection. Given the sheer number of pathogens out there, the number of different clonal lineages for B cells is estimated to be around 100 billion. A landscape like that can be difficult to navigate without a map.
Luckily, an atlas was recently published in Nature Biotechnology. It is the work of scientists collaborating between Penn’s own Perelman School of Medicine and faculty from the School of Biomedical Engineering, Science and Health Systems at our next-door neighbor, Drexel University. Using tissue samples from an organ donor network, the authors, led by Nina Luning Prak, MD, PhD, of Penn and Uri Heshberg, Ph.D., of Drexel, submitted the samples to a process called deep immune repertoire profiling to identify unique clones and clonal lineages. In total, they identified nearly a million lineages and mapped them to two networks: one in the gastointestinal tract and one that connects the blood, bone marrow, spleen, and lungs. This discovery suggests that the networks might be less complicated than initially thought. Also, it confirms a key role for the immune system in the gut.
Not only does this B cell atlas provide valuable information to the scientific community, but it also could serve as the basis for immune-based therapies for diseases. If we can identify these lineages and how clonal selection occurs, we could identify the most effective immunological cells and perhaps engineer them in the lab. At the very least, the extent to which scientists understand how B cells are formed and develop has received an enormous push with this research.
Understanding Muscle Movement
Natural movements of limbs require the coordinated activation of several muscle groups. Although the molecular composition of muscle is known, there remains a poor understanding of how these molecules coordinate their actions to confer power, strength, and endurance to muscle tissue. New fields of synthetic biology require this new knowledge to efficiently produce naturally inspired muscle substitutes.
Responding to this challenge, scientists at Carnegie Mellon University, including Philip R. LeDuc, Ph.D., William J. Brown Professor of Mechanical Engineering and Professor of Biomedical Engineering, have developed a computational system to better understand how mixtures of specific myosins affect muscle properties. Their method, published in PNAS, uses a computer model to show that mixtures of myosins will unexpectedly produce properties that are not the average of myosin molecular properties. Instead, the myosin mixtures coordinate and complement each other at the molecular level to create emergent behaviors, which lead to a robustness in how the muscle functions across a broad range. Dr. LeDuc and his colleagues then confirmed their model in lab experiments using muscle tissue from chickens. In the future, this new computational method could be used for other types of tissue, and it could prove useful in developing treatments for a variety of disorders.
Determining Brain Connectivity
How the brain forms and keeps memories is one of the greatest challenges in neuroscience. The hippocampus is a brain region considered critical for remembering sequences and events. The connections made by the hippocampus to other brain regions is considered critical for the hippocampus to integrate and remember experiences. However, this broad connectivity of the hippocampus to other brain areas raises a critical question: What connections are essential for rewiring the brain for new memories?
To offer an explanation for this question, a team of scientists in Hong Kong published a paper in PNAS in which they report on a study conducted in rats using resting-state function MRI. The study team, led by Ed X. Wu, Ph.D., of the University of Hong Kong, found that stimulation of a region deep in the hippocampus would propagate more broadly out into many areas of the cortex. The stimulation frequency affected how far this signal propagated from the hippocampus and pointed out the ability for frequency-based information signals to selectively connect the hippocampus to the rest of the brain. Altering the frequency of stimulation could affect visual function, indicating that targeted stimulation of the brain could have widespread functional effects throughout the brain.
Although human and rodent brains are obviously different, these findings from rats offer insights into how brain connectivity emerges in general. Similar studies in humans will be needed to corroborate these findings.
Seeing Inside a Tumor
Years of research have yielded the knowledge that the most effective treatments for cancer are often individualized. Knowing the genetic mutation involved in oncogenesis, for instance, can provide important information about the right drug to treat the tumor. Another important factor to know is the tumor’s chemical makeup, but far less is known about this factor due to the limitations of imaging.
However, a new study published in Nature Communications is offering some hope in this regard. In the study, scientists led by Xueding Wang, Ph.D., associate professor of biomedical engineering and radiology at the University of Michigan, used pH-sensing nanoprobes and multiwavelength photoacoustic imaging to determine tumor types in phantoms and animals. This new technology is based on the principle that cancerous cells frequently lower the pH levels in tissue, and designing probes with properties that are pH sensitive provides a method to find tumors with imaging methods and also treat these tumors.
With this technology, Dr. Wang and his colleagues were able to obtain three-dimensional images of pH levels inside of tumors. Importantly, it allowed them to noninvasively view the changes in a dye injected inside the tumor. Although a clinical application is years away, the information obtained using the Michigan team’s techniques could add significantly to our knowledge about tumorigenesis and tumor growth.
The Role of Bacteria in MS
The growing awareness of how bacteria interact with humans to affect health has led to the emergence of new scientific areas (e.g., human microbiome). Research findings from scientists collaborating between Caltech and UCSF suggest bacteria can play a role in the onset of multiple sclerosis. These investigators include Sarkis K. Mazmanian, Ph.D., Luis B. and Nelly Soux Professor of Microbiology and a faculty member in the Division of Biology and Biological Engineering at Caltech. Reporting their research results in PNAS, the researchers found several bacteria elevated in the MS microbiome. Study results showed that these bacteria regulated adaptive immune responses and helped to create a proinflammatory milieu. The identification of the bacteria interacting with immunity in MS patients could result in better diagnosis and treatment of this disabling disease.
