by Kat Sas

The availability of rapid, accessible testing was integral to overcoming the worst surges of the COVID-19 pandemic, and will be necessary to keep up with emerging variants. However, these tests come with unfortunate costs.
Polymerase chain reaction (PCR) tests, the “gold standard” for diagnostic testing, are hampered by waste. They require significant time (results can take up to a day or more) as well as specialized equipment and labor, all of which increase costs. The sophistication of PCR tests makes them harder to tweak, and therefore slower to respond to new variants. They also carry environmental impacts. For example, most biosensor tests developed to date use printed circuit boards, or PCBs, the same materials used in computers. PCBs are difficult to recycle and slow to biodegrade, using large amounts of metal, plastic and non-eco-friendly materials.
In addition, most PCR tests end up in landfills, resulting in material waste and secondary contamination. An analysis by the World Health Organization (WHO) estimated that, as of February 2022, “over 140 million test kits, with a potential to generate 2,600 tonnes of non-infectious waste (mainly plastic) and 731,000 litres of chemical waste (equivalent to one-third of an Olympic-size swimming pool) have been shipped.”
In order to balance the need for fast, affordable and accurate testing while addressing these environmental concerns, César de la Fuente, Presidential Assistant Professor in Bioengineering and Chemical and Biomolecular Engineering in the School of Engineering and Applied Science, with additional primary appointments in Psychiatry and Microbiology within the Perelman School of Medicine, has turned his attention to the urgent need for “green” testing materials.
The de la Fuente lab has been working on creative ways to create faster and cheaper testing for COVID-19 since the outbreak of the pandemic. Utilizing his lab’s focus on machine biology and the treatment of infectious disease, they created RAPID, an aptly named test that generates results in minutes with a high degree of accuracy. An even more cost-effective version, called LEAD, was created using electrodes made from graphite. A third test, called COLOR, was a low-cost optodiagnostic test printed on cotton swabs.
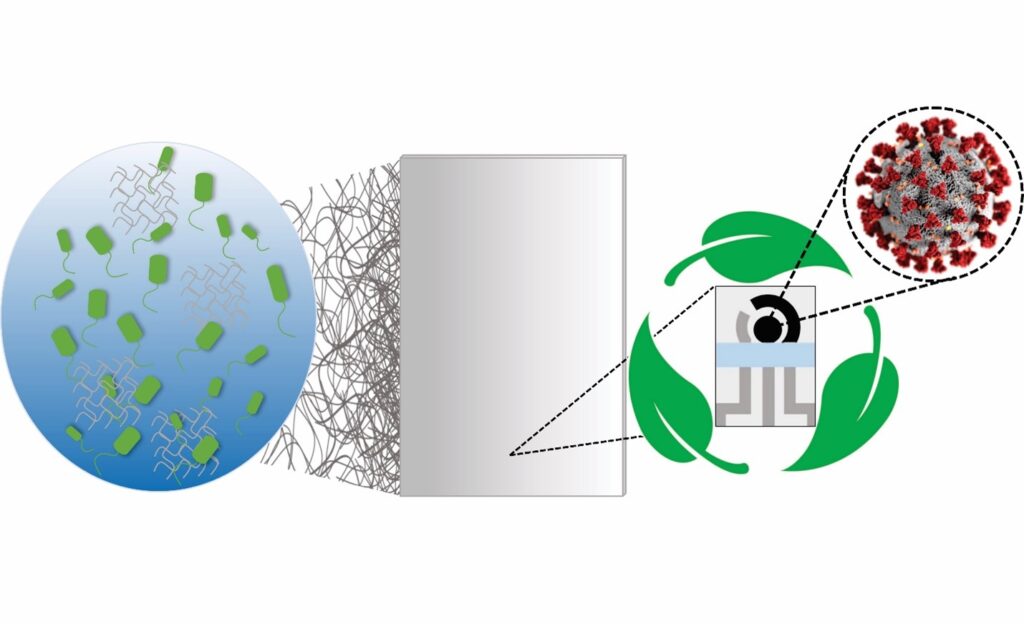
The team’s latest innovation incorporates the speed and cost-effectiveness of previous tests with eco-friendly materials. In a paper published in Cell Reports Physical Science, the group introduces a new test made from Bacterial Cellulose (BC), an organic compound synthesized from several strains of bacteria, as a substitute for PCBs.
Read the full story in Penn Engineering Today.



 Cosette Tomita, a master’s student in Bioengineering, spoke with Penn Engineering Graduate Admissions about her research in cellular therapy and her path to Penn Engineering.
Cosette Tomita, a master’s student in Bioengineering, spoke with Penn Engineering Graduate Admissions about her research in cellular therapy and her path to Penn Engineering.
 A research team led by engineers at the
A research team led by engineers at the